
Unbinding
Kinetics
beyond binding affinity
Koffee™ is our computational drug discovery technology for classical computers — delivering results faster than it takes to brew a cup of coffee, and with no quantum hardware required yet.
Koffee™ Unbinding Kinetics brings kinetics to early-stage drug discovery.
This video requires marketing cookies to be enabled.
While binding affinity is typically the primary focus in the early stages of drug discovery, protein-ligand kinetics can strongly influence efficacy, dosing, toxicity, and selectivity.
But although computational methods for binding affinity are abundant, tools for kinetics have so far been limited, with most approaches relying on computationally intensive molecular dynamics simulations. In contrast, Koffee™ Unbinding Kinetics delivers physics-based residence time estimates in just 1—2 minutes per complex on a GPU, without any machine learning or target-specific fine-tuning.
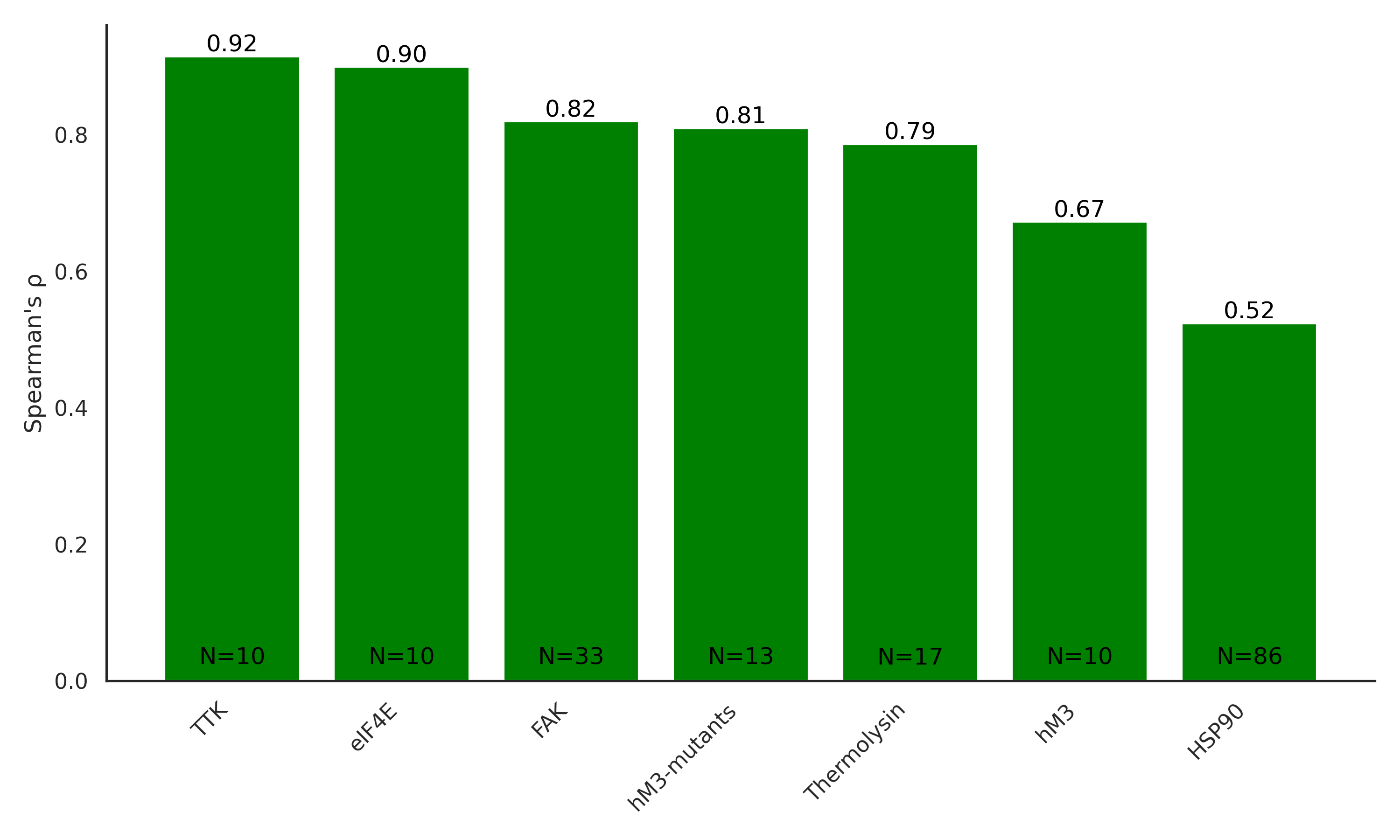
Performance on Public Data Sets
Ranking correlation of model predictions with experimental residence times from public datasets covering diverse protein targets such as kinases, GPCRs, and chaperones.
Our preprint describes a real-world prospective drug discovery project, which involved predicting both binding modes and ranking compounds by residence time. In this study, we obtained strong correlation to experimental data in a prospective exercise, supporting drug discovery project decision making in real time.
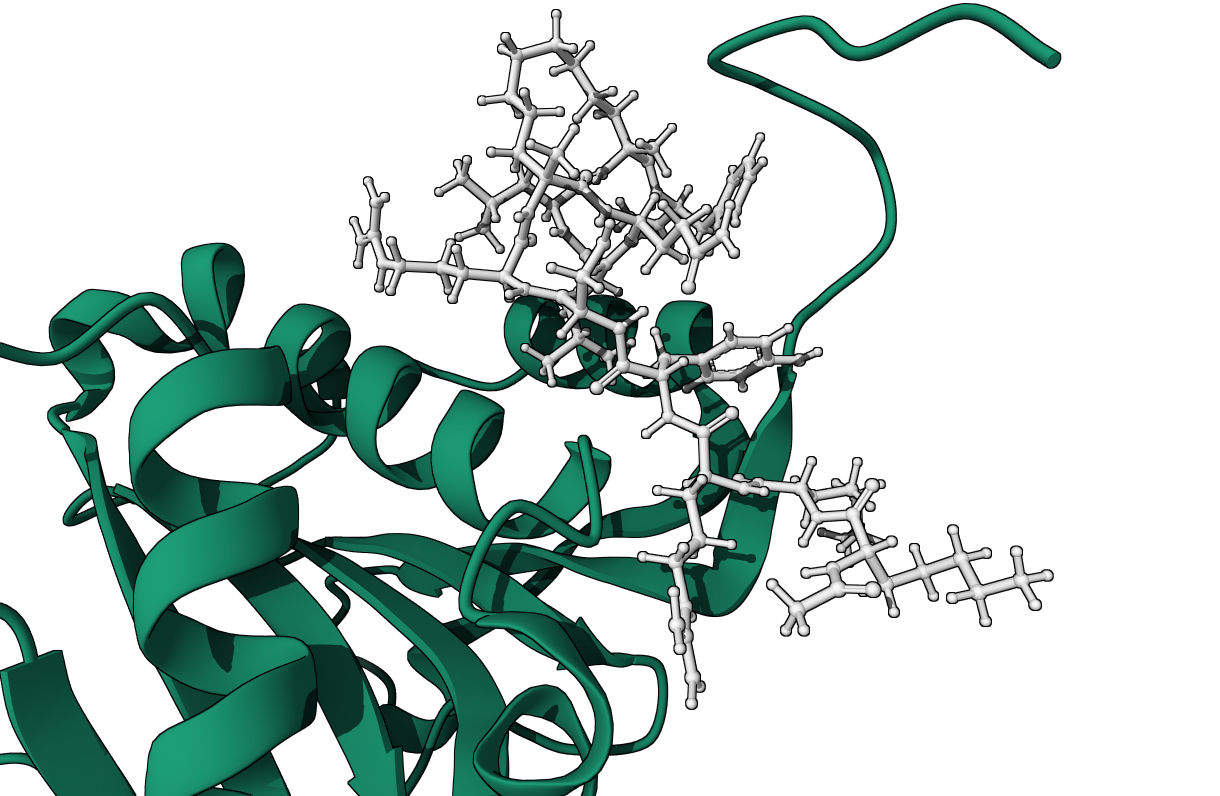
Chemical Diversity
Our approach ranks chemically diverse ligands — including those with different scaffolds and charge states — and spanning in size from fragments to large macrocycles.
In contrast, most binding affinity methods only perform reliably on congeneric series.
Use Cases
Reduce Off-Target Toxicity
Drugs that fail to bind their targets long enough may show reduced efficacy and increased side effects. Optimizing residence time helps improve on-target efficacy while minimizing off-target toxicity.
Assess the Impact of Binding Site Mutations
Compounds that retain long residence times across common resistance mutations or genotypic variants represent more robust and resilient drug candidates.
Enhance Virtual Screening with Kinetics Rescoring
Integrate kinetics into your virtual screening workflow to identify long-lasting binders in the final selection stages and improve hit quality.
Optimize PROTAC Design with Residence Time Insights
Proteolysis Targeting Chimeras (PROTACs) degrade target proteins by recruiting an E3 ligase to catalyze ubiquitination. Achieving the right residence time is crucial for effective degradation.